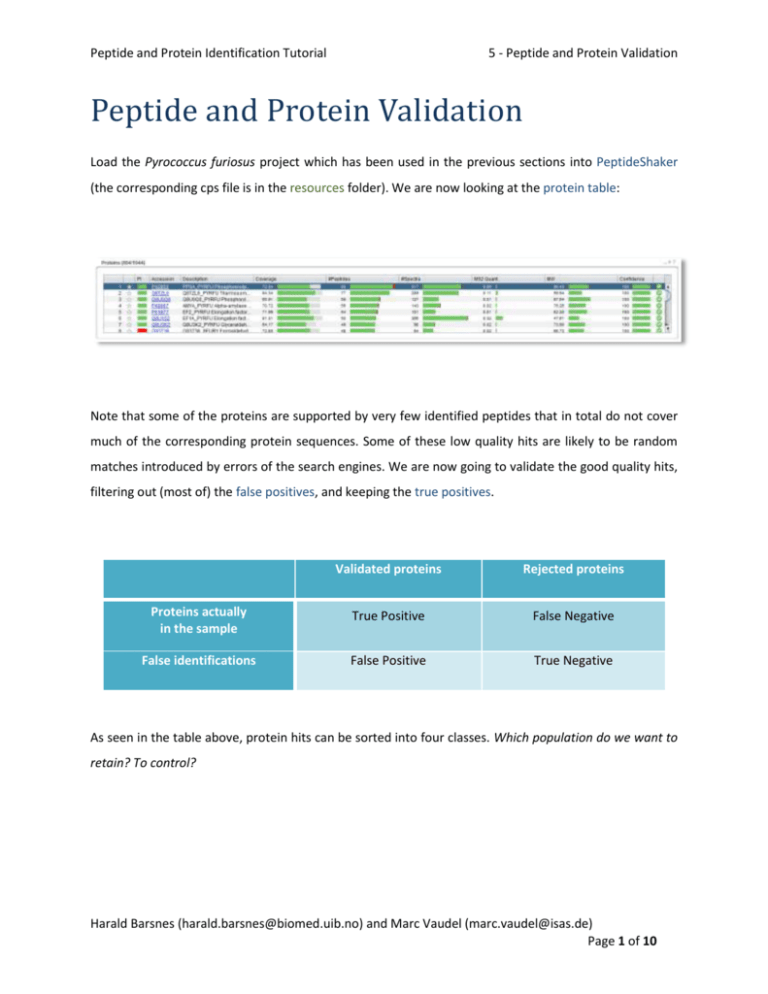
 
Systems biology enables the integration of data on molecular changes in the body in health and disease. The phenomenon of modification is accompanied, as a rule, by an increase in the area available for the solvent of the modified amino acid residue and its active environment. Results revealed that PTMs are localized in stable and compact space protein globule motifs that are exposed to a solvent. For proteins with PDB structures, a comparative analysis of the structural changes accompanying the modifications was performed. Fifteen proteins containing PTMs were identified in blood samples from patients with kidney cancer. The proteins were analyzed using ultra-high resolution HPLC-MS/MS and structural analysis was performed with the AMBER and GROMACS software packages. 
Conformational changes in proteins after modification were analyzed. We examined protein PTMs in the blood samples from patients with kidney cancer. It is likely that dysregulation of post-translational cellular signaling leads to abnormal proliferation and oncogenesis. Common PTMs are reversible and serve as a mechanism for modulating metabolic trans-formations in cells. Mascot search engine in PD was conducted due to the higher compatibility with the data compared to PeptideShaker and Peaks.Post-translational modification (PTM) leads to conformational changes in protein structure, modulates the biological function of proteins, and, consequently, changes the signature of metabolic transformations and the immune response in the body. The labeling of samples was determined to be conducted before enrichment due to low yield when labeling after enrichment. The procedure was validated using HeLa cells, which worked successfully. The method conducts the micro column protocol, which has proven to be more successful than the batch mode protocol. The labeling step was determined to be after enrichment based on the number of confident identification (147 phosphopeptides labeled before enrichment versus 3 phosphopeptides labeled after enrichment).Ĭonclusion: In this this study, a method for quantifying the phospho proteome in adipose tissue has been presented. The procedure was validated using HeLa cell with 1068 phosphopeptides confidently identified. PhosSTOP had a negative effect with the results and was therefore excluded from further experiments, as the phosphopeptides identified with PhosSTOP were lower and less confident. Micro column protocol was more successful in terms of number of identification (252) compared to the batch mode protocol (147 identified). PD was also successful in interpreting the fragment spectra giving relatively good information about the fragments compared to Peaks and PeptideShaker. Mascot was unique in 51 phosphopeptides compared to Peaks’ 33 unique phosphopeptide identifications. Results and discussion: Mascot in PD identified the highest number of phosphopeptides with 252 confident identifications compared to Sequest HT (146), Peaks (239) and PeptideShaker (197). The post-processing of data output was done on PD, Peaks and PeptideShaker to determine the most compatible search engine. The last step in procedure was to determine the labeling of the samples (before or after enrichment). The next step was to validate the procedure by running the procedure on HeLa cells. to determine which protocol yielded the highest number of phosphopeptides. The samples were enriched with TiO2 using the enrichment protocols published by Dickhut et al. Materials and methods: The approach employed in this thesis is a bottom-up based method. In this thesis, present a novel bottom-up method for the quantitative analysis of the phospho proteome in adipose tissue. Quantitative analysis of the proteome with tandem mass tag is a technique for calculating the relative abundance of the proteome in tissues and organelles. Background: Mass spectrometry-based proteomics has increasingly been the choice of method for the global analysis of the composition, modification and dynamics of proteins. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
